Chapter 4 Taxonomy Profiling
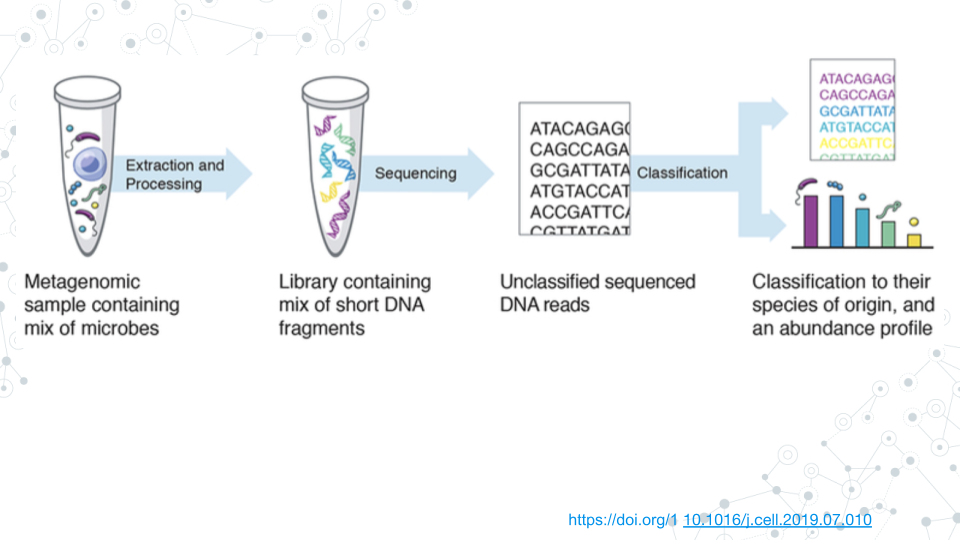
Taxonomic profiling is the process of assigning metagenomics sequencing reads to taxa (e.g., genus, species) and estimating the relative abundance of organisms in a sample. Due to the high complexity of metagenomics data, long-read shotgun sequencing offers many advantages over short-read sequencing for taxonomic classification.
In this module, we will work with long-read metagenomic data and learn to classify and visualize taxonomy on the Galaxy platform. We will use Kraken 2 to assign taxonomy to sequencing reads and Krona to visualize results interactively.
This Taxonomy Profiling module offers hands-on experience with real data. We will
- Learn core concepts through lecture material including taxa, taxonomic classification, and species abundance.
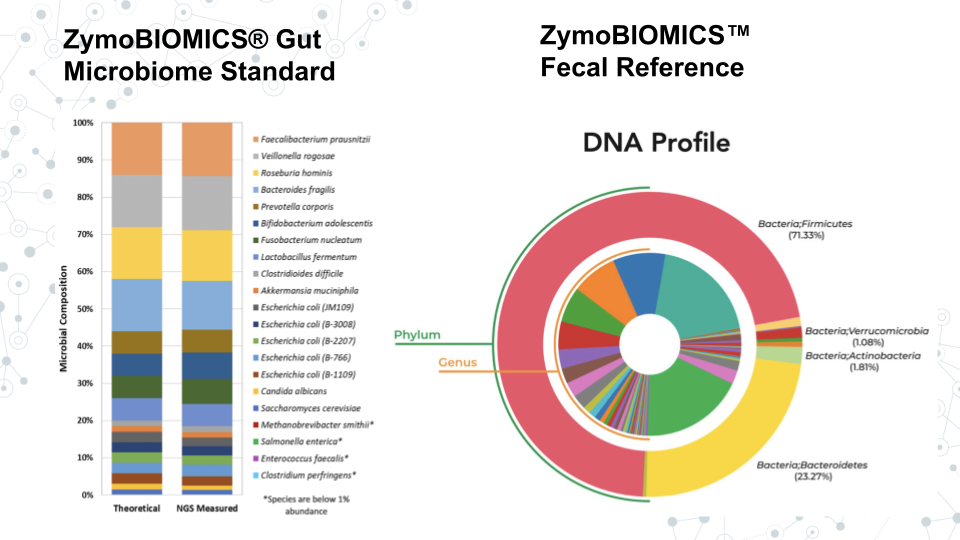
- Activity: Explore taxonomic profiles from the Zymo Research gut microbiome standard using output of Kraken 2. This standard contains a known mixture of microbes with expected relative abundances. You will examine the structure of the taxonomy output and perform some basic analyses.
- Prelab: Run taxonomy profiling ourselves in Galaxy using raw sequences from the gut microbiome standard to reproduce and interpret the results using Kraken 2 for classification and Krona pie charts for visualization.
- Project: Taxonomically classify a soil metagenome sample (collected by the BioDIGS initiative) which serves as an example of an uncurated, potentially novel data. We will compare the soil results with the gut standard to learn what to expect when working with real-world samples.